GEMSiRV User Manual is provided in Manual.pdf.
The main features of GEMSiRV are summarized below.
Reconstruction
Click on Reconstruction in the menu bar to open Model databases and Reference databases.
- Model importing and editing (Step-by-step)
Right click on Model databases to Import SBML file (.xml) or to Import spreadsheets (.xls), you can import a metabolic model in SBML/spreadsheet format as described in Metabolic Models.
You can directly edit/update the content of the imported model.
- Reference database construction (Step-by-step)
Right click on Reference databases to Import database (.xls), you can import a reference database provided in Reference Databases to construct your own reference database.
You can add the information about metabolites and reactions described in a model to the reference database by right clicking on the metabolic model to add rxn&met to the Ref. DB.
- Draft reconstruction (Step-by-step)
A draft reconstruction can be generated by mapping a blank reconstruction output by GBKParser, containing gene information only, to a reference reconstruction. You just simply right click on the blank reconstruction to Draft a reconstruction
Given a user-provided list of orthologous genes, GEMSiRV will extract from the reference model of the "orthologous reaction" (i.e. reactions that are catalyzed by the protein products of orthologous genes) that conform to the gene-protein-reaction (GPR) associations defined in the reference model.
- Model refinement (Step-by-step)
You can right click on a model to Generate simulation tables, so that a mathematical model including a stoichiometric matrix which describes the connectivity feature of the network as well as default systems boundaries can be generated.
With the generated simulation tables, you can identify dead-end metabolites.
Then, you can right click on the model with simulation tables to Define environmental conditions, e.g. the complete medium to simulate all extracellular metabolites can enter/exit the cell freely (complete medium.TXT), and the in silico (computational) minimal media for the model E. coli textbook (M9 medium comp.TXT).
Simulation
Click on Simulation in the menu bar to perform the analyses implemented in GEMSiRV including
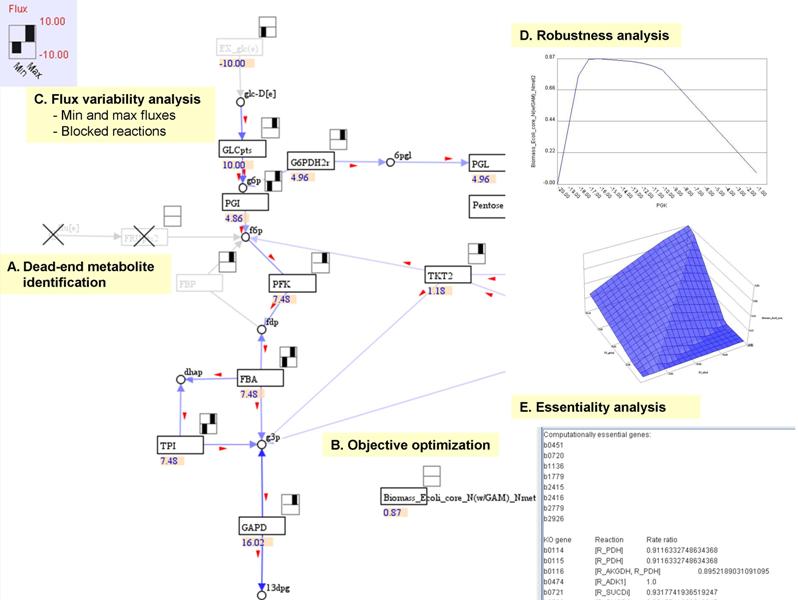
- Dead-end metabolite identification (Step-by-step)
Select a metabolic model and a map (if you have), GEMSiRV can identify dead-end metabolites and tag them with crosses in the map.
- Objective optimization (Step-by-step)
Select a metabolic model and a map (if you have) for objective optimization, the flux result can be visulized in the map.
- Flux variability analysis (Step-by-step)
Select a metabolic model and a map (if you have) for flux variability analysis, the min and max fluxes of reaction can be plotted in the map and the blocked reaction are tagged with crosses.
- Robustness analysis (Step-by-step)
Select the reactions of interest in a model to see how sensitive the objective is to the particular reactions.
- Essentiality analysis (Step-by-step)
Select a metabolic model for essentiality analysis, the computational essential genes or reactions can be identified.
- Gene deletion analysis (Step-by-step)
Select a metabolic model for gene deletion analysis, the gene-deletion model can be saved and imported for further evaluation.
Visualization
Click on Visualization in the menu bar to open Maps wherein you can create metabolic maps.
- Metabolic map creaction (Step-by-step)
Right click on Maps to Create new map. You can add maps, reactions, metaboltes and lines to create a map onto the main network view window.
- KEGG map loading (Step-by-step)
Right click on Maps to Load KEGG maps by either Import KEGG map (.xml) or Retrieve KEGG map.
- Map replacement (Step-by-step)
With two lists for metabolite and reaction mapping, you can right click on a map to Replace caption of nodes to convert the map to a customized map. KEGG to Model SEED and KEEG to MetaCyc mapping lists are provided here (met KEEGtoSEED.TXT, rxn KEGGtoSEED.TXT, met KEEGtoMetaCyc.TXT and rxn KEEGtoMetaCyc.TXT), but the mapping lists are not guaranteed to completely correct.
- Information extraction (Step-by-step)
Right click on a map to Extract reaction information from a model and choose a model you want to extract information from. Then you can show the extra information of reaction in the map by right clicking a reaction to Show extra info..
- Flux visualization (Step-by-step)
Right click on a map to Load reaction fluxes, the reaction fluxes were be laid on the reactions in the map. A file containing flux simulations of aerobic and anaerobic conditions in the model of Out_textbook.xml is provided here (flux withorwithout o2.TXT).
- Gene expression visualization (Step-by-step)
Right click on a map to Load gene expressions, the gene expressions were be superimposed around the associated reactions in the map. An expression profiling of oxygen effects on E. coli is provided here (Expr o2 effect on Ecoli rep.TXT).